Case study – Ethanol-guided hybridisation of extracellular vesicles with liquid-crystalline lipid nanoparticles
SPARTA® provides single-particle, label-free compositional confirmation of true EV–LCNP hybrid formation
Researchers at the University of Florence developed a one-step, ethanol-guided microfluidic method to fuse red-blood-cell-derived extracellular vesicles (RBCEVs) with liquid-crystalline lipid nanoparticles (LCNPs) to generate cubosome hybrid nanoparticles. Using structural, biochemical and single-particle analyses, including label-free SPARTA® Raman analysis, they confirmed the formation of true hybrids that retained both the LCNP cubic structure and the desired beneficial RBCEV properties.
The challenge
Red-blood-cell-derived extracellular vesicles (RBCEVs) are membrane-bound nanoparticles that are nonimmunogenic. They retain native lipids and membrane proteins that enhance biocompatibility, prolong circulation time and support efficient cellular uptake, making them attractive drug delivery vehicles. However, EVs can be difficult to load efficiently and are challenging to manufacture at scale.
Liquid-crystalline lipid nanoparticles (LCNPs), specifically cubosomes, have a bicontinuous cubic internal structure which provides high cargo-loading capacity, a large internal surface area and membrane curvature that can improve drug delivery inside cells. But LCNPs lack the biocompatibility and targeting properties of EVs.
Hybridising RBCEVs with LCNPs therefore offers a way to combine the biological advantages of EVs with the structural and loading benefits of LCNPs. However, achieving this balance has proven difficult.
Common fusion techniques such as freeze–thaw cycles, extrusion and sonication can damage EV membranes or strip away important proteins, while gentler co-incubation approaches often cause the LCNP cubic structure to collapse, removing the very features that make them effective at carrying and delivering drugs.
To overcome these limitations, researchers at the University of Florence developed an ethanol-guided microfluidic strategy that brings the two systems together as the nanoparticles are forming. Instead of trying to force already-made particles to fuse, the hybrids are created in a single step under carefully controlled conditions, allowing LCNPs to assemble and incorporate EV components at the same time.
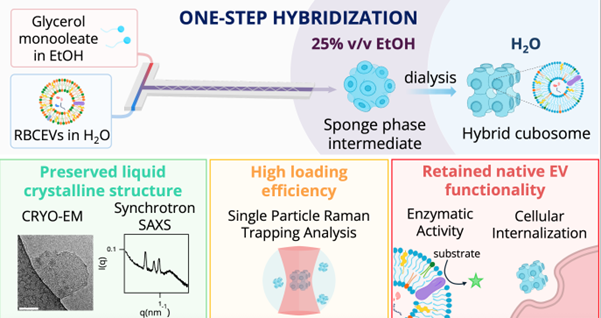
In this study, they set out to determine whether this approach could successfully generate true hybrid nanoparticles that keep the LCNP cubic structure intact, successfully include RBCEV components, and preserve the activity of key proteins.
Proving that this had occurred, however, is analytically challenging because traditional techniques have several limitations:
- Bulk measurements: These average signals across large particle populations, masking particle-to-particle differences and making it difficult to determine how uniformly hybridisation has occurred.
- Size and morphology-focused methods: Many techniques can reveal particle size, shape or internal structure, but provide little information about the chemical composition of individual nanoparticles. This makes it challenging to confirm whether biological components are truly incorporated within each particle.
- Labelling-based approaches: Fluorescent or chemical labels can alter native membrane behaviour or interfere with protein function, and often cannot distinguish surface attachment from genuine membrane fusion.
The solution
To overcome these analytical limitations, the researchers incorporated single particle automated Raman trapping analysis (SPARTA®) into their workflow.
SPARTA® measures nanoparticles at the single-particle level by combining optical trapping with Raman spectroscopy to record the chemical fingerprint of individual particles. This makes it possible to see if both lipid and EV-derived protein signals are present within the same nanoparticle.
Importantly, SPARTA® is label-free and non-destructive. It avoids fluorescent or chemical modification that can alter membrane structure or protein behaviour, while still providing detailed compositional information that structural methods alone cannot deliver.
Approach
In this study, SPARTA® was used alongside structural and biochemical methods to confirm whether the ethanol-guided microfluidic strategy was effective at producing true cubosome hybrid nanoparticles.
These included:
- Synchrotron SAXS: To determine internal phase symmetry and monitor structural transitions, assessing whether the cubic liquid-crystalline architecture was preserved.
- Cryo-electron microscopy (cryo-EM): To visualise nanoparticle morphology and internal organisation at the single-particle level.
- Western blotting: To confirm the presence of key RBCEV membrane and luminal proteins in the hybrid particles.
- Enzymatic assays: To assess retention of acetylcholinesterase activity following hybridisation.
- Cellular uptake studies: To evaluate whether incorporation of RBCEV components influenced nanoparticle internalisation.
SPARTA® added the missing information by showing by providing detailed insight into nanoparticle composition. Around 200 nanoparticles per sample were analysed in solution. Their Raman spectra were examined for characteristic lipid peaks from a key component of LCNPs, glyceryl monooleate (GMO), alongside RBCEV-associated protein signals. By determining whether both signatures were present within the same particle, the researchers could confirm hybrid nanoparticle formation and quantify the proportion of particles that were hybrids.
Outcomes
The combined structural, compositional and functional analyses showed that the ethanol-guided microfluidic strategy successfully generated structurally defined hybrid nanoparticles.
Key outcomes of the study included:
- Confirmed molecular-level hybridisation: Single-particle SPARTA® analysis demonstrated that individual nanoparticles contained both GMO lipid signals and RBCEV-associated protein peaks. This confirmed true hybrid formation.
- Preserved cubic structure: The internal cubic architecture of the LCNPs was retained following RBCEV incorporation.
- Quantified hybridisation efficiency: By analysing nanoparticles individually, SPARTA® enabled quantification of how uniformly RBCEV components were incorporated across the population, revealing the extent of particle-to-particle variability.
- Retention of EV proteins: Confirmed the presence of key RBCEV proteins within the hybrid nanoparticles.

