Case Study – Single-particle Raman analysis of extracellular vesicles as breast cancer biomarkers

A minimally invasive, more accurate approach to breast cancer detection using Raman spectroscopy
Breast cancer diagnosis currently relies on invasive biopsies and visual assessments that are not always accurate. Extracellular vesicles (EVs) are biomarkers that offer a promising non-invasive alternative, as they carry biochemical signatures that reflect their cell of origin. However, detailed, single particle analysis of their lipid, protein and nucleic acid content/composition is difficult with traditional methods. In this study, SPARTA® enabled high-throughput, single-particle Raman analysis of EVs from different breast cell types, accurately distinguishing cancer and non-cancer EVs and highlighting its potential for minimally invasive, bias-free diagnosis.
The challenge
Breast cancer is the most common cancer affecting women worldwide, with more than two million new cases diagnosed in 2020. For effective treatment selection and the best prognosis, fast and accurate diagnosis essential.
However, current diagnostic approaches rely on physical examination, imaging, and tissue biopsy with histopathological analysis, which are invasive, prone to sampling error, and dependent on individual interpretation, leading to variability between clinicians and laboratories. These limitations mean there is an urgent need for less invasive diagnostic approaches that reduce interpretation variability and bias.
EVs are emerging as a promising diagnostic biomarker in this space. These nanoscale particles are released by cells and carry different combinations of proteins, lipids, and nucleic acids that reflect their cell of origin. By analysing these molecular differences, it is possible to distinguish cancer-derived EVs from non-cancerous ones. This makes EVs a valuable tool for cancer detection and monitoring using minimally invasive blood samples.
Despite their potential, there are still challenges to address in using EVs for diagnostics. These include:
- Sample interference from labelling and immobilisation
Many EV analysis methods rely on labelling or immobilisation steps that increase sample interference and introduce artefacts. These manipulations can alter the native physical and biological properties of EVs, generate background signal, and amplify contaminating particles, reducing confidence that measured signals truly reflect EV composition. - Bulk analytical methods don’t capture heterogeneity
Many conventional methods analyse EVs in bulk, averaging signals across highly heterogeneous populations. This masks biologically and diagnostically relevant EV subpopulations and obscures subtle molecular differences that may be critical for accurate disease detection. As a result, current approaches often fail to capture the full molecular complexity and diversity of EVs. - EVs are difficult to purify from other particles
EVs are hard to isolate cleanly from blood because they exist alongside many other tiny biological particles, particularly lipoproteins, that are similar in size and behaviour. As a result, bulk analysis methods often measure a mixture of particles rather than EVs alone, making it difficult to know where a given molecular signal is really coming from. Analysing EVs at the single-particle level is therefore essential to separate true EV subpopulations from contaminating particles and avoid misleading results.
The solution: Raman spectroscopy using SPARTA®
Raman spectroscopy provides a promising way to address the limitations of existing EV analysis approaches. As a label-free, non-destructive light-based technique, it enables direct measurement of biological samples without chemical modification, preserving EVs in their native state. It can also be automated, reducing user involvement and interpretation bias.
In this study from Imperial College London, Raman spectroscopy was implemented using our Single Particle Automated Raman Trapping Analysis (SPARTA®) platform to investigate the potential of EVs as biomarkers of breast cancer. While Raman spectroscopy has previously been applied to cancer cells and tissue, these approaches typically rely on invasive biopsy procedures and are unsuitable for routine monitoring. Here, Raman analysis was instead applied to EVs isolated from blood samples, enabling a minimally invasive approach.
SPARTA® overcomes key limitations of conventional EV analysis by enabling single-particle analysis of EVs. This makes it possible to capture EV heterogeneity, resolve distinct subpopulations, and distinguish EV-derived molecular signals from background or contaminating particles, all of which would be masked with bulk methods.
Approach
In this study, EVs were isolated from 11 breast cell lines, including 2 non-cancerous and 9 cancerous types, and analysed individually using SPARTA® to capture their Raman spectra. Each spectrum provided a chemical fingerprint reflecting the relative contributions of key biomolecular components such as proteins, lipids and nucleic acids.
More than 14,000 individual EVs were measured across multiple cell types, producing large, standardised datasets for compositional analysis.
Raman spectra from SPARTA® were analysed using a multivariate classification model to distinguish between cancer-derived and non-cancer-derived EVs. The model was trained on EVs from a first isolation and validated using a second, independent isolation to assess reproducibility.
This analysis revealed consistent differences in EV chemical composition between the two groups, with variations in biomolecular make-up, particularly the relative contributions of lipids and proteins, emerging as the main drivers of discrimination.
Each individual EV was assigned a quantitative score based on its chemical profile, enabling clear and reproducible separation of cancerous and non-cancerous EV populations across isolations.
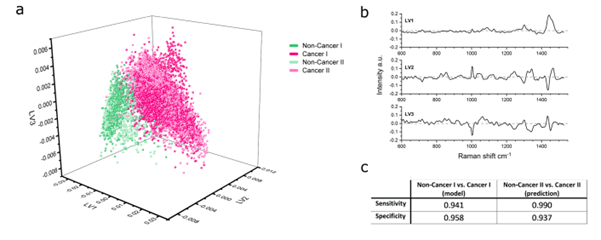
Fig 1: Multivariate analysis of Raman spectra from SPARTA® enables discrimination between cancer-derived and non-cancer-derived EVs.
Outcome
Using this approach, SPARTA® was able to distinguish between cancer-derived and non-cancer-derived EVs with very high performance, achieving close to 99% sensitivity and over 95% specificity. These results highlight the potential of SPARTA® as an unbiased, reliable, label-free method for identifying cancer-associated EVs based on their molecular composition.
Following this, SPARTA® spectra were used to explore the underlying differences in EV composition in more detail. This analysis focused on how key components, such as nucleic acids, lipids and proteins, varied across EVs from different cell types. This revealed clear compositional differences between cancerous and non-cancerous EVs, helping to explain why the two groups could be distinguished so accurately and offering deeper insight into EV biology.

