Case study – Coupling lipid nanoparticle structure and single-particle composition analysis to design enzyme-responsive nanocarriers

SPARTA® reveals how LNP composition and structure influence intracellular processing and function
Lipid nanoparticles (LNPs) are important in modern drug delivery, but rationally designing stimuli responsive particles is challenging. In this study, researchers from Imperial College London and the Karolinska Institute designed LNPs that respond to the enzyme phospholipase D (PLD). By combining structural and single-particle compositional analysis using SPARTA®, they were able to observe how PLD interacts with individual LNPs in real-time, and understand how their composition and structure influence enzyme responsiveness. These insights open new possibilities for the rational design of enzyme-responsive nanocarriers and controlled drug release.
The challenge
Lipid nanoparticles are highly versatile delivery systems with tuneable physicochemical properties that make them ideally suited for vaccines and RNA therapeutics. However, rationally designing LNPs that respond predictably to stimuli is limited by a lack of understanding of how nanoparticle composition and structure are linked to intracellular processing and function.
In this study, researchers were interested in looking at LNPs that are responsive to the enzyme PLD, which is known to be dysregulated in diseases including neurodegeneration, vascular disease and some cancers.
PLD modifies certain lipids in the nanoparticle membrane, converting them into a negatively charged lipid called phosphatidic acid (PA). This chemical change can reorganise the membrane and alter its structure, potentially making the nanoparticle more permeable or less stable. This means PLD activity could be used as a biological trigger for controlled drug release.
However, until now, it has been unclear how PLD interacts with LNPs, limiting efforts to design particles that deliberately exploit this enzyme.
Progress in this area has been limited by several analytical challenges that mean it is not possible to perform dynamic reaction monitoring to effectively study stimuli responsiveness:
- Bulk methods mask particle-to-particle differences
Most techniques analyse LNPs in bulk, averaging signals across heterogeneous populations and hiding how individual nanoparticles respond to enzymatic activity. - Structural techniques lack compositional detail
Techniques such as small-angle X-ray and neutron scattering (SAXS/SANS) provide detailed structural information but cannot resolve particle-to-particle differences or be used to study dynamic reaction monitoring. - Label-based methods can interfere with nanoparticle behaviour
Many techniques rely on labels to track nanoparticle changes, but these can alter membrane properties, interfere with enzyme activity, or obscure true chemical changes.
The solution
This study combines Single Particle Automated Raman Trapping Analysis (SPARTA®) with SAXS/SANS to link lipid nanoparticle composition, internal structure and morphology, and to monitor dynamic interactions between LNPs and PLD.
SPARTA® is a label-free, non-destructive technique that analyses individual nanoparticles using Raman spectroscopy, enabling researchers to measure subtle differences in lipid composition at the single-particle level. This made it possible to follow enzyme-driven changes in real time, revealing how phospholipase D (PLD) interacts with and alters individual nanoparticles, and how these effects vary from particle to particle.
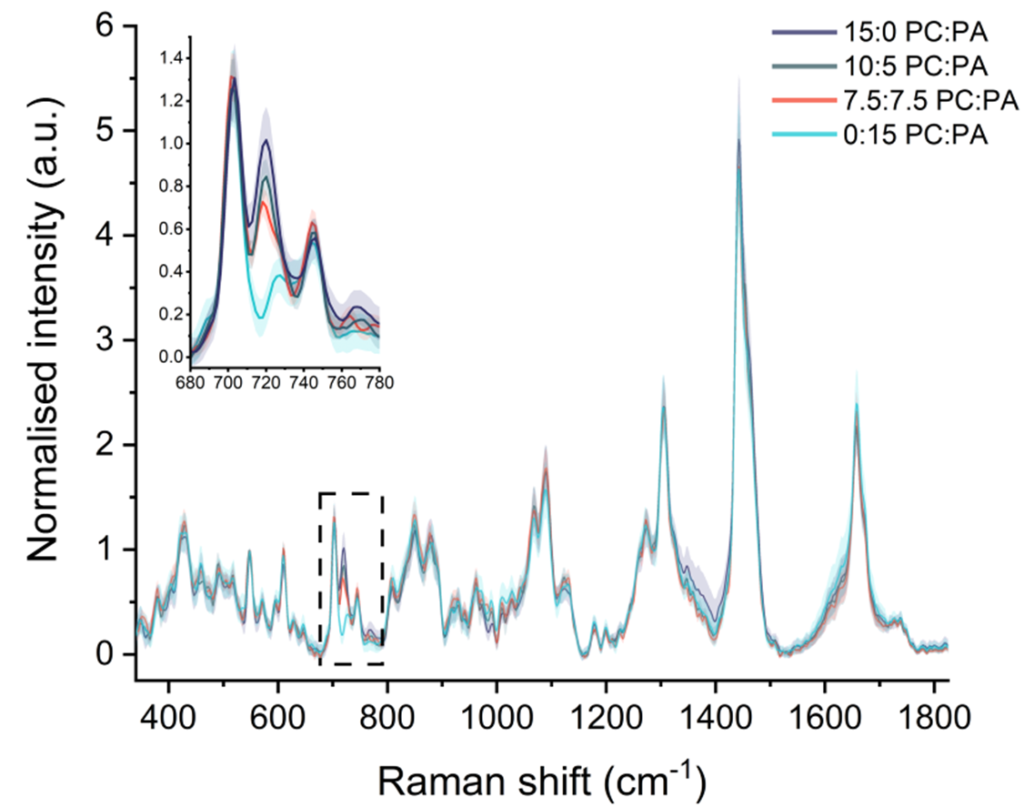
Fig 1: Raman spectra from individual lipid nanoparticles show how SPARTA® can detect subtle differences in lipid composition at the single-particle level.1
SAXS/SANS were used to examine the internal structure of lipid nanoparticles, showing how enzyme-driven changes in lipid composition led to changes in internal architecture. Used alongside SPARTA®, they linked chemical changes to physical structure, helping explain how LNP architecture influences enzyme interactions.
Approach
The researchers created LNPs with a complex internal structure and included phosphatidylcholine (PC), a lipid that PLD can act on. These nanoparticles were then exposed to different amounts of PLD, as well as an inactive form of the enzyme for comparison.
Using SPARTA®, the team analysed one nanoparticle at a time, measuring each particle repeatedly to see how its chemical composition changed over time. Tracking signals linked to the PC headgroup, the researchers could directly measure how much PC was converted into PA inside individual nanoparticles.
By repeating this process across hundreds of nanoparticles, they were able to see both overall trends across the population and differences between individual particles. Changing the amount of PLD controlled how quickly the reaction happened.
At the same time, SAXS and SANS showed how increasing PA levels and PLD activity reshaped internal water channels, membrane curvature and overall nanoparticle morphology.
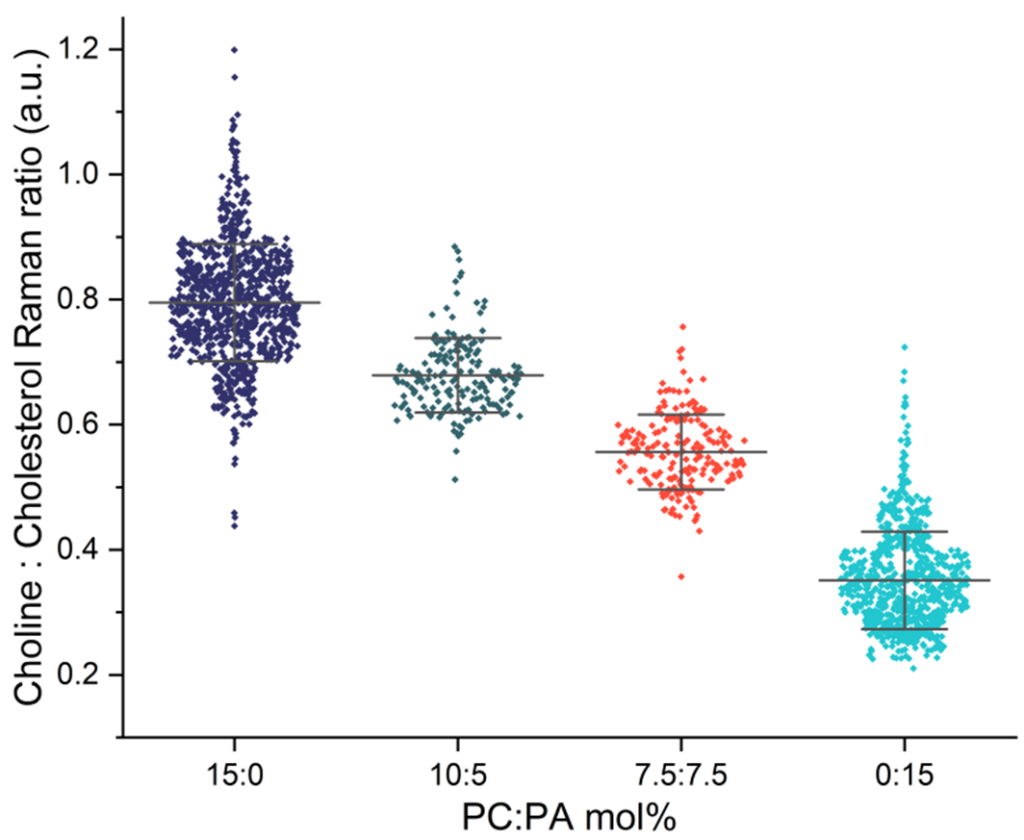
Fig 2: Comparison of choline signals from individual lipid nanoparticles, highlighting compositional differences across LNP formulations.
Outcome
This combined approach overcame key analytical challenges by enabling label-free, real-time monitoring of chemical changes in LNPs at the single-particle level. It allowed researchers to:
- Observe how enzymes act on individual nanoparticles as reactions unfold, rather than relying on bulk measurements that average behaviour and hide important particle-to-particle differences.
- Track genuine chemical changes without using labels, avoiding interference with enzyme activity or nanoparticle structure and providing a more accurate picture of how LNPs respond to biological stimuli.
The study revealed several important insights:
- PLD can fully access and remodel the entire LNP membrane, converting neutral lipids into negatively charged ones far more efficiently than in comparable vesicles.
- PLD activity is strongly influenced by LNP membrane structure and curvature.
- Structural tuning of the LNP membrane directly influences this enzymatic activity, highlighting the potential to design enzyme-responsive LNPs.
These findings point to new strategies for disease-responsive payload delivery in diseases where PLD is dysregulated. With the help of SPARTA®’s unique compositional insight, this study also helps address a key challenge in LNP design by linking composition, structure and biological function at the single-particle level.

